|
12/26/2023 0 Comments Postview software 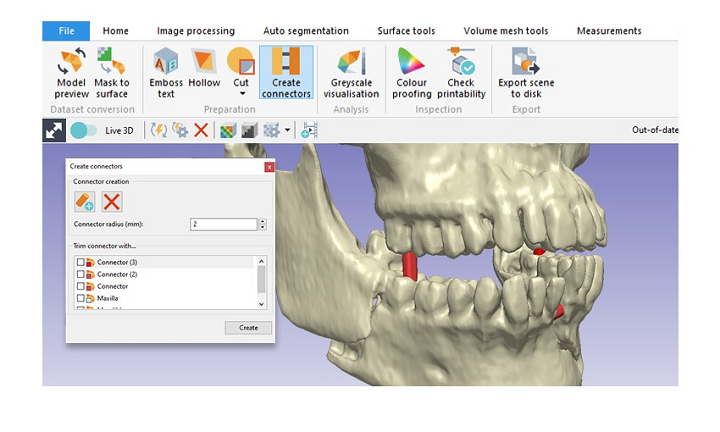
Let’s compare it to backbone (using coffeescript in the following examples). ItemView is typically initialized with a model and is shown in the DOM. Let’s see how we can apply each of these to improve our code. While backbone only has the Backbone.View class, marionette extends this class with 4 types of specialized views: ItemView, CollectionView, CompositeView and Layout. Views are the most abused component of backbone apps, where the bulk of ugly/boilerplate code gets thrown in. This means that many of its components extend backbone classes, which makes refactoring incredibly easy. The best thing about marionette is that it builds on top of backbone. In this article, I will focus on how to refactor a backbone application using marionette.js -a framework that can mitigate many of backbone’s shortcomings. With a steep learning curve, shortage of opinionated patterns, lots of boilerplate code, and poor memory management/cleanup strategies, backbone usage can be discouraging. In this article, I will focus on how to refactor a backbone application using marionette.js -a framework that can mitigate many of backbone’s shortcomings.ĭespite its popularity, backbone.js has many drawbacks. From Modeling to Medicinal Chemistry: Automatic Generation of Two-Dimensional Complex Diagrams.Despite its popularity, backbone.js has many drawbacks. (1996) A fast flexible docking method using an incremental construction algorithm, Journal of Molecular Biology, 261, 470-489. Journal of Chemical Information and Computer Sciences, 44, 1065-1078. (2004) Automated Generation of Structural Molecular Formulas under Constraints. Fricker, P., Gastreich, M., and Rarey, M. (2006) Molecular Complexes at a Glance: Automated Generation of two-dimensional Complex Diagrams. Hydrogen bonds can be labeled by their energy or length. Interacting amino acids can be displayed as structure diagrams of complete molecules or of the interacting molecule part. Complex diagrams can be shown in a browser or written to a file in one of the following formats: PNG, PDF, SVG or EPS. The selection of the shown amino acids is based on the interaction calculations which is done by the FlexX-library.Ĭomplex of Estrogen Receptor Alpha and Diethylstilbesterol. The coordinates of the structure diagram and their layout modifications are generated by the library SDGlib. Hydrophobic contacts between the ligand and protein are represented by spline segments which mark hydrophobic regions of the ligand and the corresponding amino acid descriptors of the PDB-database. 
In a complex diagram hydrogen bonds are represented as dashed lines and both the interacting amino acids of the receptor and the ligand are represented as structure diagram. The tool PoseView automatically generates two-dimensional diagrams of complexes with known three dimensional structures while adhering to conventions of chemical structure diagram generation.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
